Background:
Clinical studies often rely on extracting structured data points from free-text medical documentation—a time-consuming and error-prone process. To streamline this workflow, we developed Raki, an open-access platform (https://www.raki-data.net) that facilitates Large-Language-Model (LLM) assisted data extraction with integrated text anonymization and structured variable definitions. The goal of this work was to evaluate both the time efficiency and the extraction accuracy of Raki with respect to manual corrections needed during the human-in-the-loop process.
Methods:
We evaluated Raki using data from 188 anonymized pulmonary hypertension outpatient encounters, including diagnosis lists, written ECG reports, and spiroergometry summaries. Datapoints were extracted manually and using the Raki workflow with GPT-4o-mini hosted in the EU and human-in-the-loop correction. Data processing was performed on anonymized data in the EU and therefore, GDPR compliant. Human expert labels served as the ground truth. Extraction times were recorded using a stopwatch protocol. Uncorrected Model output was compared to corrected datapoints after human in the loop to assess variable-level accuracy, F1-score, precision, and recall.
Results:
Raki-assisted extraction substantially reduced annotation time, achieving 2.17× faster extraction for diagnoses, 3.50× for ECG, and 1.49× for spiroergometry compared to manual review.
Across variables, the uncorrected data extraction showed results, with cumulated F1-scores of 0.910 (diagnoses), 0.946 (ECG), and 0.880 (spiroergometry), although variability was present.
Conclusion:
Raki enables fast and reliable datapoint extraction from text-based clinical documentation. In retrospective study workflows, Raki-assisted extraction is 1.5–3.5× faster than unassisted manual extraction while showing high per-variable extraction accuracy, making it a practical solution for clinical research data curation at scale.
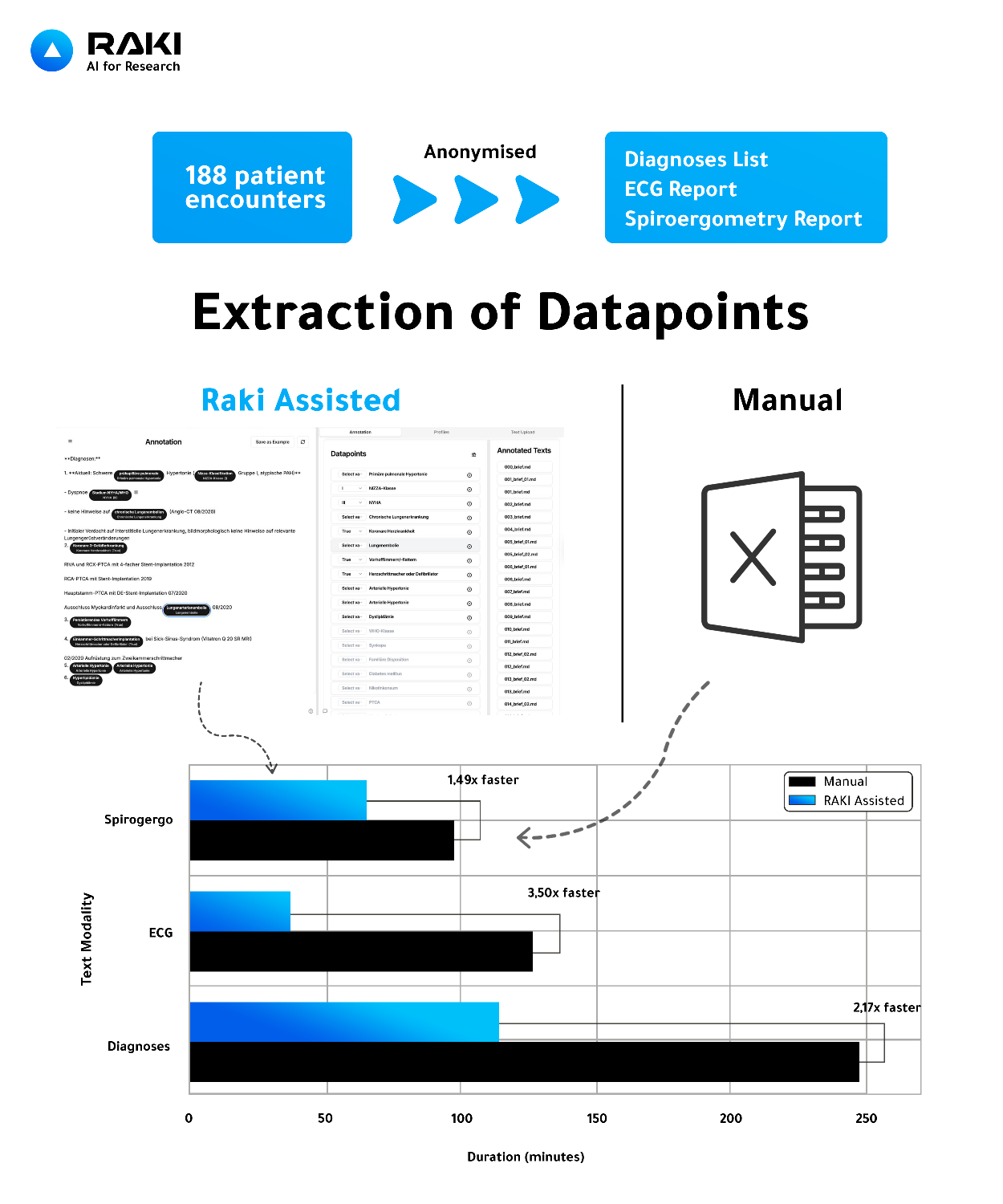
Figure 1: Comparison of manual versus Raki-assisted datapoint extraction for retrospective clinical studies. From 188 anonymized cardiology encounters, structured datapoints were extracted from diagnoses, ECG, and spiroergometry reports. Raki-assisted extraction accelerated data curation by 1.49–3.50× relative to manual review.
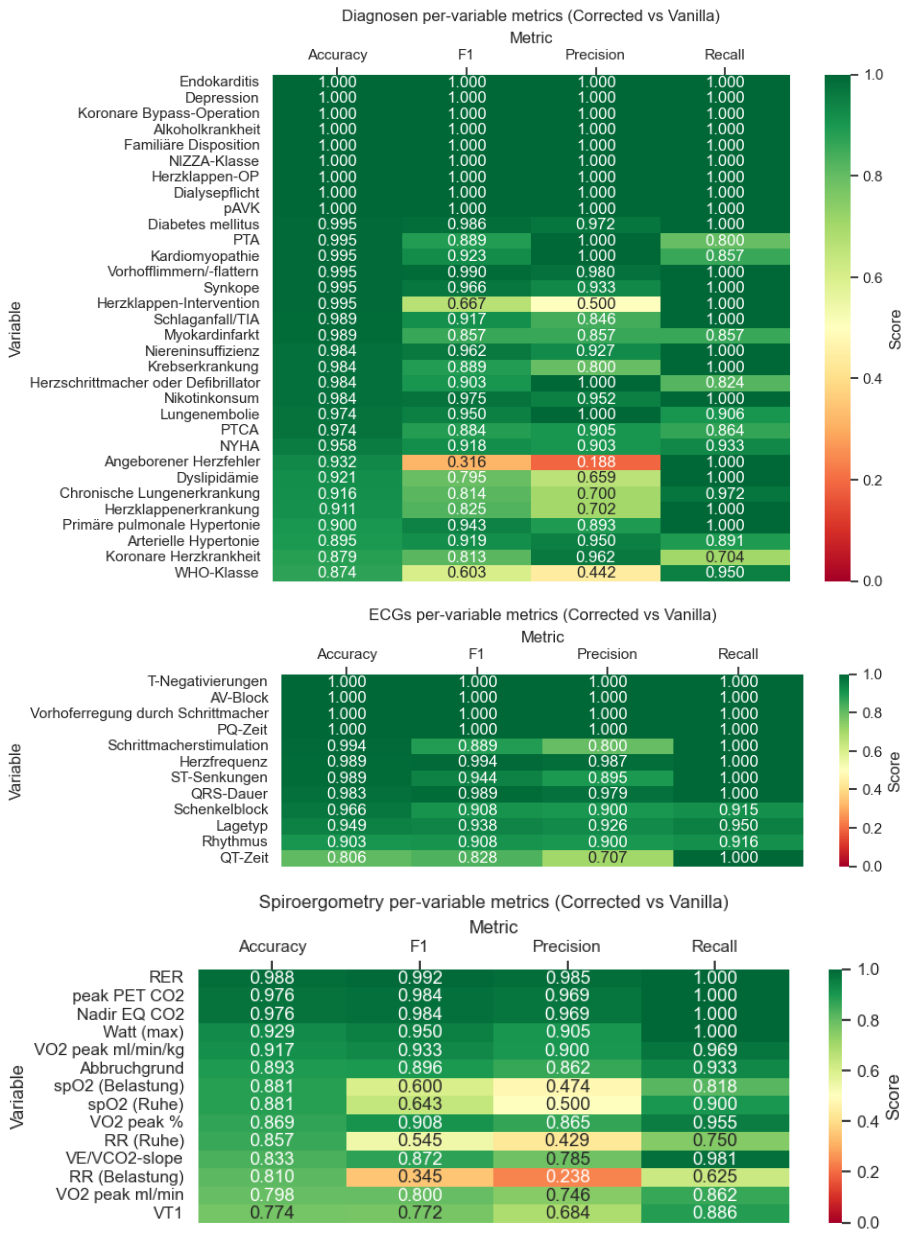
Figure 2: Per-variable comparison of uncorrected (vanilla) GPT-4o-mini output vs. human-in-the-loop corrected datapoints within the Raki platform. Accuracy, F1-score, precision, and recall are shown for each extracted variables across diagnoses, ECG, and spiroergometry reports.
