Background: Hypertrophic cardiomyopathy (HCM), the most prevalent inherited cardiac disease, is marked by striking phenotypic and clinical heterogeneity. Although impaired myocardial energetics are recognized as a hallmark of HCM, the influence of individual metabolic phenotypes on disease expression remains poorly understood. This multicenter study aimed to define and characterize patient-specific myocardial bioenergetic profiles across the HCM disease spectrum and to explore their relationships with genotype and disease severity.
Methods: Quantitative mass spectrometry-based proteomics was performed on 165 myocardial biopsies, including 118 samples from HCM patients (30% female, median age 52 [18-74] years) and 47 non-diseased donor hearts. Expression levels of 304 metabolic enzymes were integrated into a validated kinetic model encompassing 296 biochemical reactions across all major energy-generating pathways. This computational framework enabled personalized simulations of myocardial ATP generation under varying workload and substrate conditions, which were then correlated with genetic, echocardiographic, and clinical data.
Results: Compared with controls, HCM patients demonstrated a markedly reduced maximal ATP production capacity, with substantial interindividual variability (Figure 1A). Maximal uptake capacities for fatty acids and branched-chain amino acids were decreased, accompanied by a metabolic shift toward greater glucose utilization both at rest and during peak workload (p<0.01). Sarcomere mutation carriers had a 10.1 ± 4.7% lower ATP production capacity than genotype-negative HCM patients (Figure 1B). Among sarcomere-positive cases, MYBPC3 and MYH7 variant carriers showed more pronounced energetic impairment than those with other pathogenic variants (Figure 1C). HCM patients receiving beta blocker therapy exhibited a higher simulated ATP production capacity (p<0.01), associated with a higher respiratory efficacy (ATP yield per mole O2) in sarcomere-positive but not sarcomere-negative subjects.
Conclusion: This study provides a comprehensive, patient-specific map of myocardial bioenergetics, revealing distinct metabolic signatures linked to genotype and disease severity in a large, well-characterized HCM cohort. Personalized assessment of myocardial metabolism may deepen understanding of disease heterogeneity and inform precision strategies for risk stratification and targeted management in HCM.
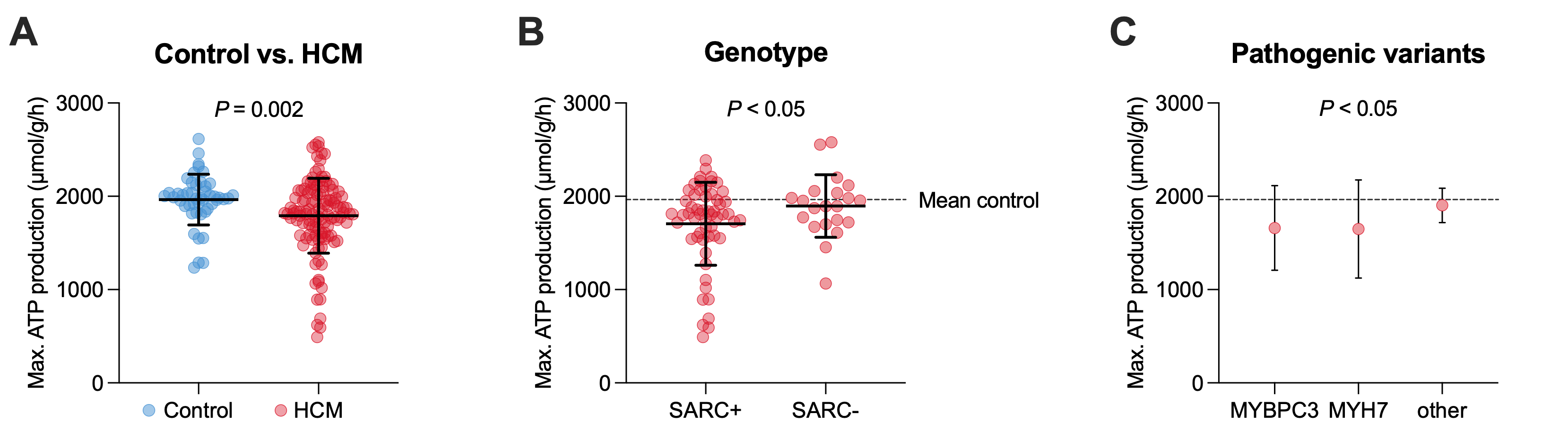
Figure 1: Patient-specific reconstruction of maximal ATP production capacity stratified by (A) disease group, (B) genotype across HCM patients, and (C) pathogenic variant in SARC+ HCM. MYBPC3, myosin-binding protein C; MYH7, myosin heavy chain 7; SARC+, sarcomere-positive; SARC-, sarcomere-negative.
