Background: Peripartum cardiomyopathy (PPCM) and dilated cardiomyopathy (DCM) share several overlapping features, yet their underlying metabolic determinants remain poorly defined. We hypothesized that integrative proteomics-based computational modeling could delineate disease-specific energetic defects in PPCM compared with DCM hearts.
Methods: High-resolution mass spectrometry-based quantitative proteomics was performed on left ventricular biopsies obtained from patients with PPCM (n=6), DCM (n=6), and non-failing controls (n=18). Protein abundance profiles of 304 distinct metabolic enzymes were integrated into a comprehensive kinetic model comprising 296 biochemical reactions governing myocardial energy metabolism. Personalized predictions of ATP production capacity, substrate utilization, and oxygen demand were performed simulating dynamic changes in myocardial workload and circulating substrate availability.
Results: Computational modeling revealed a marked reduction in maximal ATP production capacity in PPCM compared with both DCM and control myocardium (Figure 1A), whereas respiratory efficiency (ATP yield per mole O₂) was comparable. The maximal myocardial uptake capacity for fatty acids was lowered in PPCM (Figure 1B). On the molecular level, this reduced uptake capacity was linked to a PPCM-specific downregulation of key enzymes of fatty acid metabolism including long-chain-fatty-acid-CoA ligases and mitochondrial carnitine-acylcarnitine translocase (p for both <0.01). Substrate flux analysis indicated a larger contribution of lactate at peak myocardial workload in PPCM when compared to DCM and controls (16.5±2.2% vs. 11.3±5.2 vs. 10.3±7.5, respectively; Figure 1C).
Conclusions: Proteomics-informed metabolic modeling uncovers severe and disease-specific energetic failure in PPCM, exceeding that observed in DCM. The findings provide the first systems-level evidence that perturbations of myocardial metabolism and mitochondrial dysfunction may underpin PPCM, highlighting novel targets for therapeutic interventions in maternal heart failure.
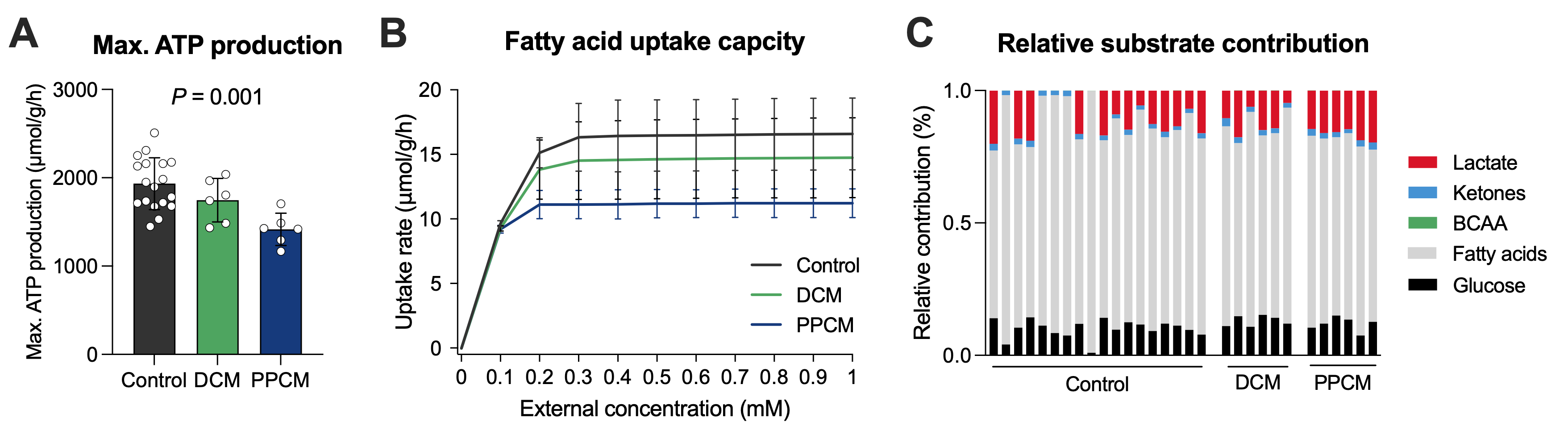
Figure 1: Computational modeling results of (A) maximal ATP production capacity, (B) myocardial uptake capacity of fatty acids, and relative contribution of energy-delivering substrates to ATP production at peak workload. Statistical significance was determined by one-way ANOVA. BCAA, branched-chain amino acids
