Introduction: High-sensitivity C-reactive protein (hsCRP) is an established biomarker recommended for cardiovascular risk stratification. Elevated hsCRP levels have been associated with adverse outcomes in large clinical studies. Conventionally, the lowest hsCRP value is considered a patient’s “true” value, while thresholds such as >2 mg/L define elevated risk. Interventional trials (e.g., with ziltivekimab) use similar cutoffs to select patients for anti-inflammatory therapy. In real-world settings, however, hsCRP levels may vary considerably due to comorbidities, medication, and other factors. This study aimed to quantify the real-world variability of hsCRP in a large cardiology cohort.
Methods: In this retrospective multicentre study, pseudonymised cardiological patient-level data, including hsCRP measurements, were extracted from the data management systems at both sites of the University Heart Centre Freiburg – Bad Krozingen via the local data integration centre. Measurements were performed using the Cardiac C-Reactive Protein (Latex) High Sensitive assay on a Cobas Analyzer (Roche, Germany). Statistical analyses were conducted in R 4.4.3. The study was approved by the local ethics committee.
Results: After applying a strict hsCRP cutoff of ≤10 mg/l the dataset contained 9,110 patients with 15,197 hsCRP measurements. Mean hsCRP was 2.07 mg/L (median 1.30 mg/L, SD 1.96, coefficient of variation (CV) 0.95). Restricting to patients with ≥2 hsCRP measurements yielded 2,585 patients with a total of 9,060 measurements. Mean hsCRP was slightly lower at 1.98 mg/L (median 1.55 mg/L) with substantial intra-individual variability (SD 1.54, CV 0.78). In the restricted cohort median age was 71 years (IQR 20) and 70% were male (measurement period December 2010 until July 2024). Median time between measurements was 185 days.
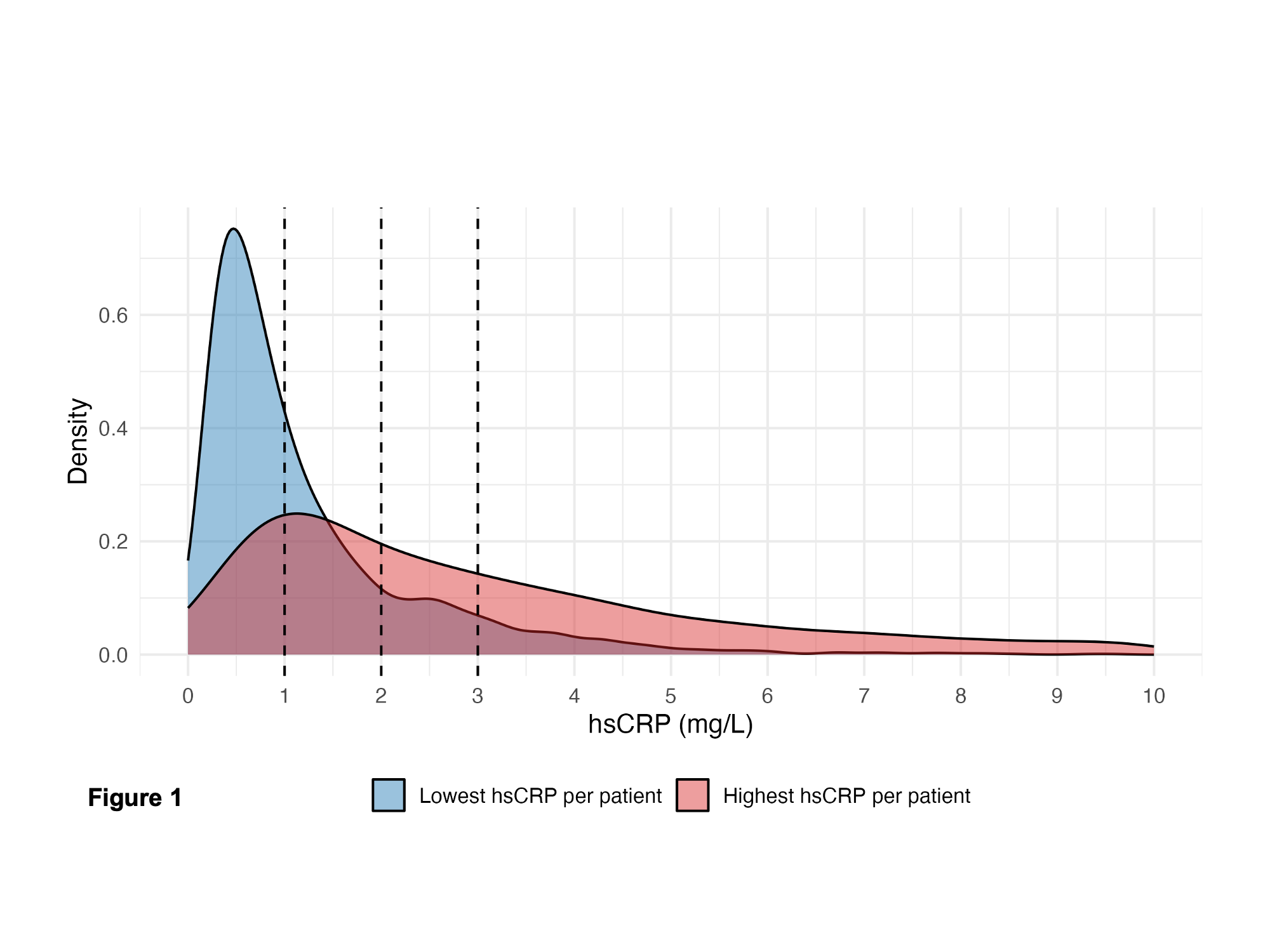
Figure 1 demonstrates patient-individual highest and lowest hsCRP levels in patients with hsCRP ≤10 mg/l and ≥2 tests. Assuming an individual’s lowest hsCRP value represents their true level, we calculated that 38% of patients would have been misclassified as ≥2 mg/L if their highest value had been used instead.
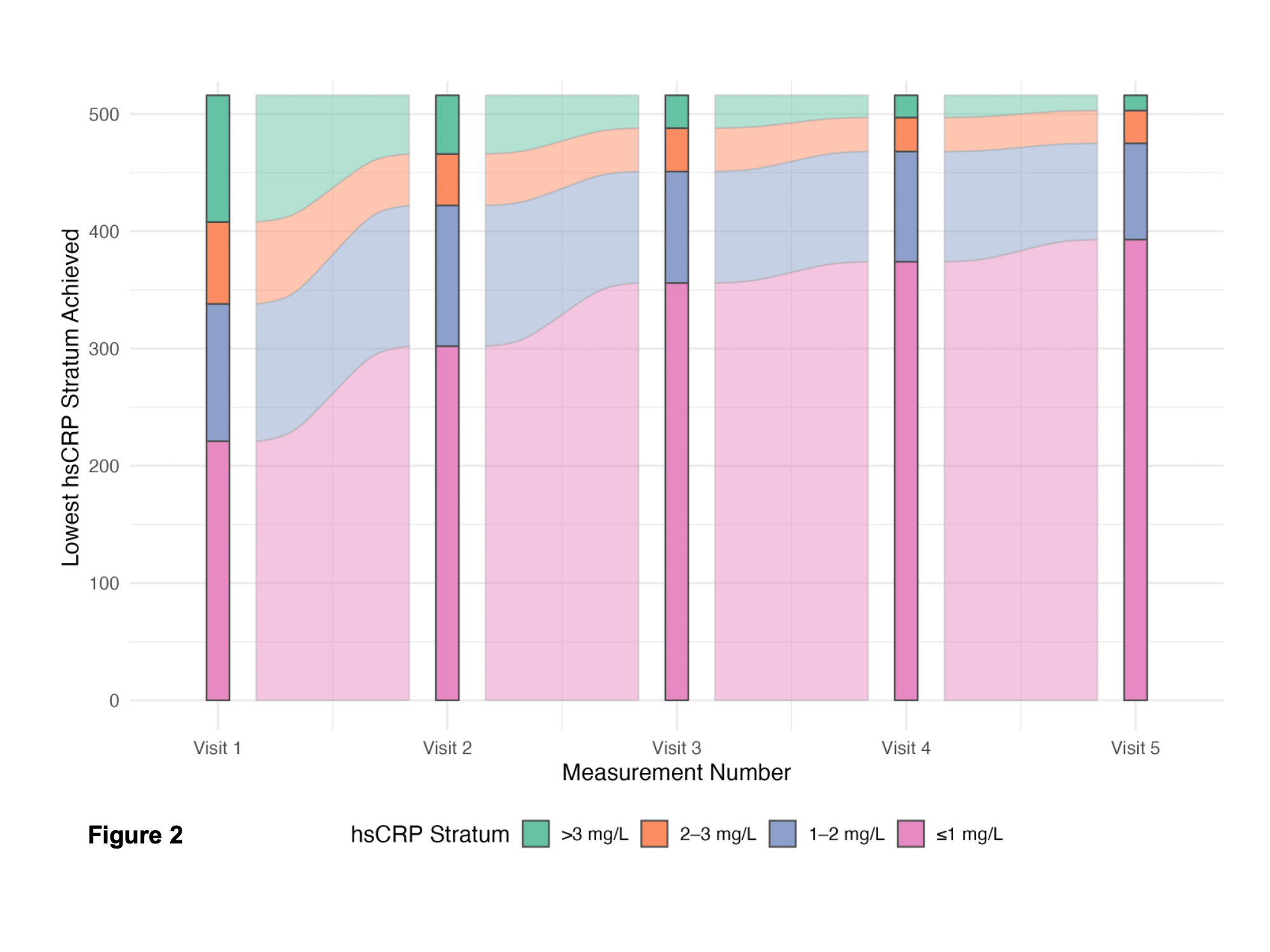
To determine how serial testing would influence patient risk categorization we restricted our analysis to a subset of patients with ≥5 measurements and hsCRP ≤20 mg/l (Figure 2). When reaching their lowest value (considered "true"), they remained in that stratum even if subsequent measurements yielded higher values. Median time between measurements was 250 days. This analysis yielded a progressive increase of the ≤1mg/l stratum (from 43% at first visit to 72% after 5 visits).
Conclusion: HsCRP levels show pronounced intra-individual variability even within the “non-inflammatory” range (≤10 mg/L). A single measurement may thus misclassify a substantial proportion of patients into higher risk categories. Serial testing could improve risk stratification accuracy. These findings highlight the need to account for real-world hsCRP variability when basing clinical or therapeutic decisions on this biomarker.