Background:
Large language models (LLMs) show promise for extracting structured data from text-based medical documentation. We developed “Raki” (https://raki-data.net/), a platform hosted in the European Union that imports clinical reports, extracts and anonymizes their text, and enables automated data extraction using LLMs. This study aims to measure the performance of data extraction using the platform across different LLMS when applied to real-world reports, comparing their performance against manually validated reference data.
Methods:
ECG, Spiroergometry, Echocardiography and diagnoses reports from 188 pulmonary hypertension patients were processed through Raki. All text content was extracted from our clinical information systems and anonymized before analysis in Raki. In total, 84 distinct diagnostic and quantitative parameters were annotated, yielding 53,144 value-level comparisons. Four LLMs (GPT-5-mini, GPT-5-nano, GPT-4o-mini, DeepSeek-Chat) were benchmarked using standardized prompts. Outputs were structured in JSON format. Model performance was evaluated against expert-labeled ground truth using precision, recall, F1-score, and accuracy. All patient data were fully anonymized in accordance with GDPR, ensuring no identifiable information was processed.
Results:
All models showed high performance across domains (F1: 0.915–0.988). GPT-5-mini achieved the most consistent results (e.g., diagnoses F1 = 0.988, ECG F1 = 0.984), while DeepSeek-Chat delivered highest precision for echocardiography (0.934). Structured endpoints such as diagnoses reached >97% accuracy. More variable content like spiroergometry showed larger performance differences across models. Average error rates ranged from 0.4 to 5.4 per 100 comparisons, depending on domain and model.
Conclusion:
This study presents a detailed evaluation of LLMs applied to real-world clinical documentation using the novelly developed Raki platform. Our results show consistent and reliable high quality data extraction across different state of the art LLMs. Our results confirm the potential of automated datapoint extraction for retrospective clinical studies using LLMS. While this study analyzed pulmonary hypertension patients, Raki could be employed for other specialties and text domains as well.
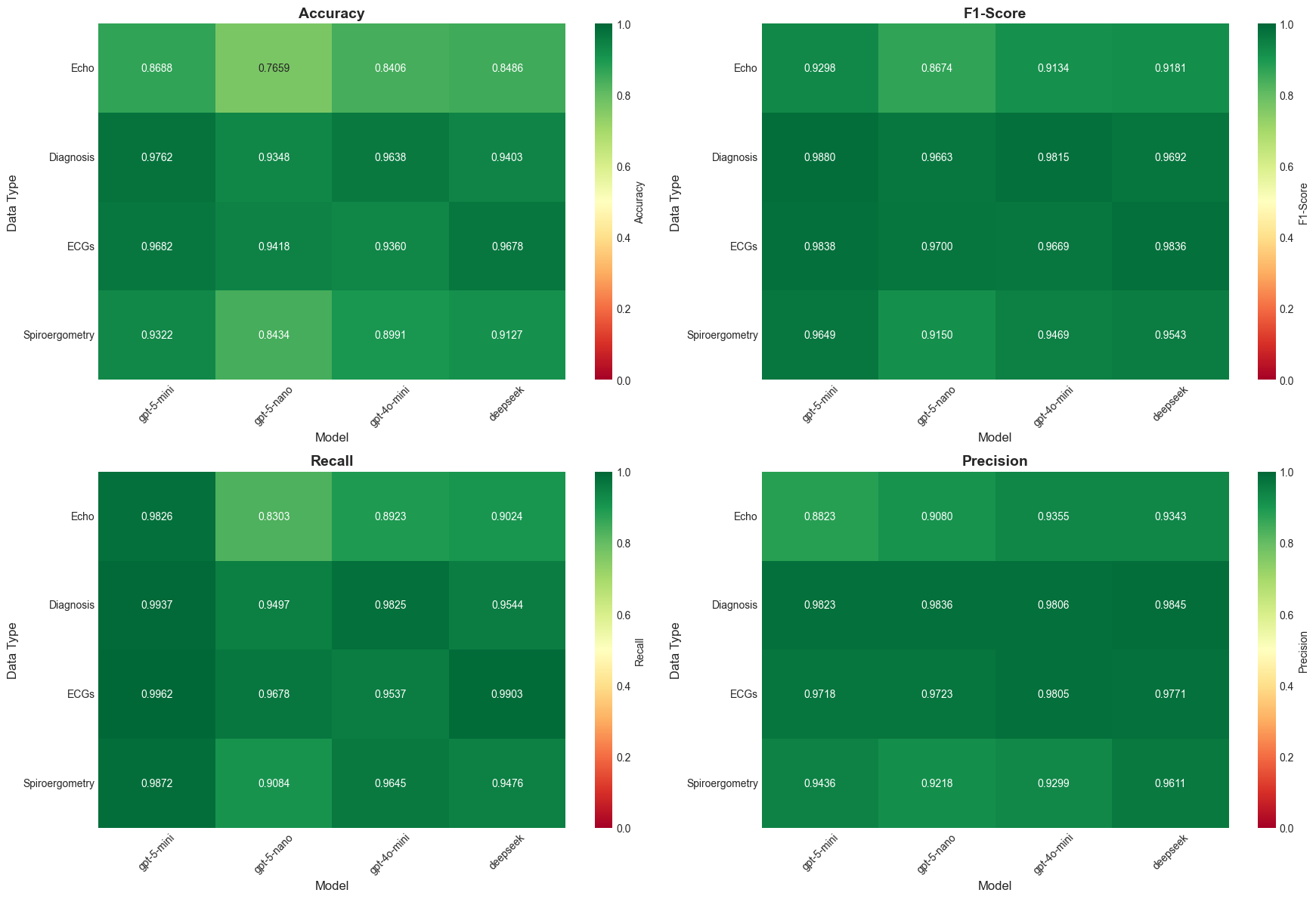
Figure 1. Heatmaps display Accuracy, F1-Score, Recall, and Precision across models and text domains (Echo, Diagnoses, ECGs, Spiroergometry). Color intensity reflects performance (red: low, green: high).
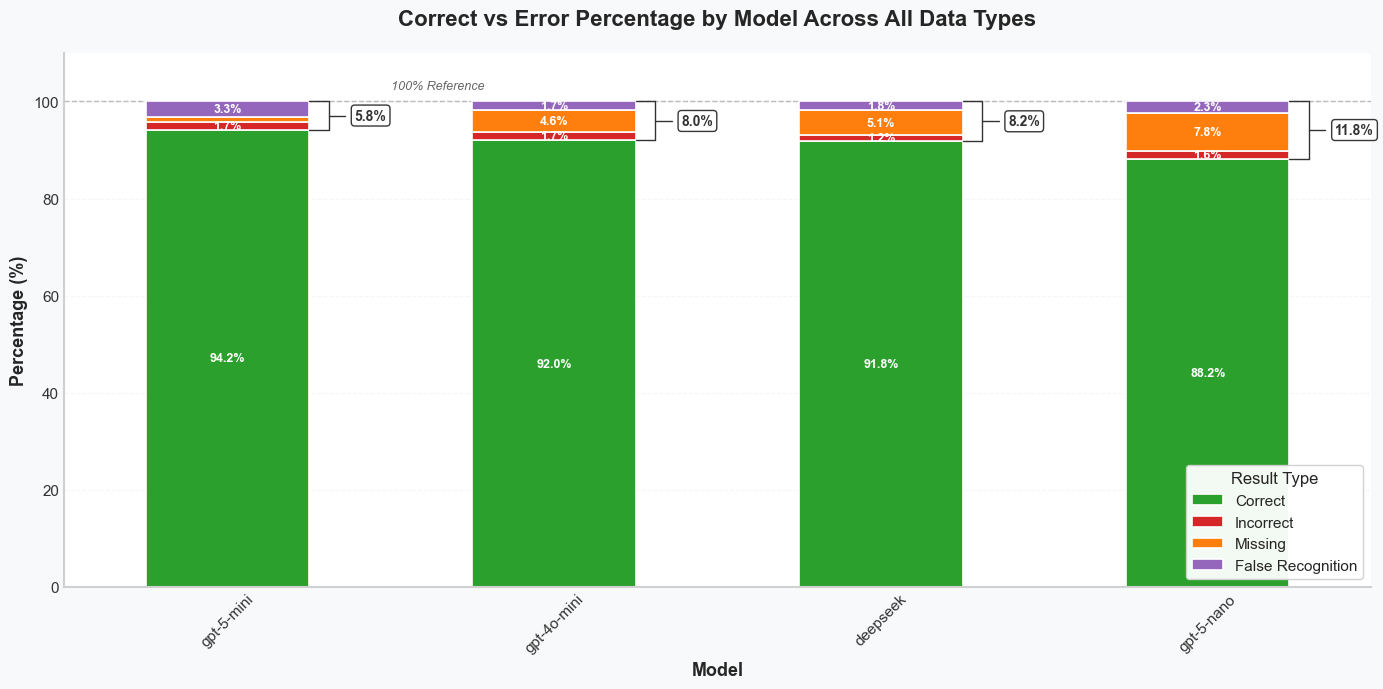
Figure 2. Stacked bars show correct vs. error rates (Incorrect, Missing, False Recognition) per model across all domains. Green indicates correct predictions; colored segments represent error types.
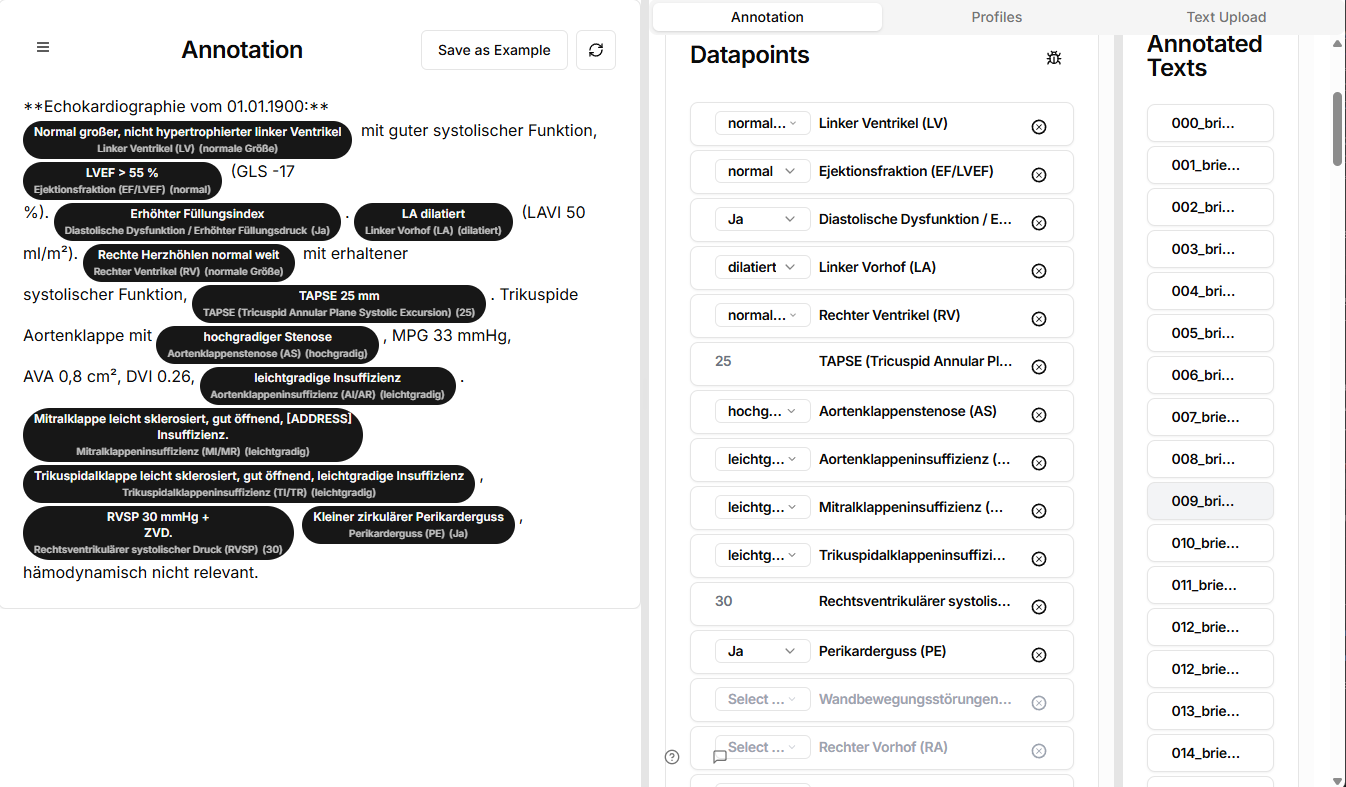
RAKI platform interface post-anonymization. Reports are parsed into structured datapoints for benchmarking LLM outputs against expert-validated ground truth.
Profiles sample in excel file
